Image-Generation
cr_mech_coli contains extensive methods to render realistic microscopic images synthetically. We are not only able to estimate the parameters of our computational model via our Fitting-Methods, but can also optimize the parameters of the image generation pipeline in order to completely clone existing microscopic images. This unique combination means that we can fully embedd our model and data-generation pipeline into real world scenarios. The crm_gen module achieves this and also provides a script crm_gen which can be used from the command line and thus does not require and programmatic experience with the used methods.
In essence, our pipeline functions by utilizing the vertices of our Model and then generates 3D surfaces with pyvista [3] which are then projected along the z-axis. Afterwards, a series of filters is applied which capture effects such as environmental noise, halo effects surrounding cells or other optical transformations such as diffraction.
The following images showcase the functionality of the imaging capabilities.
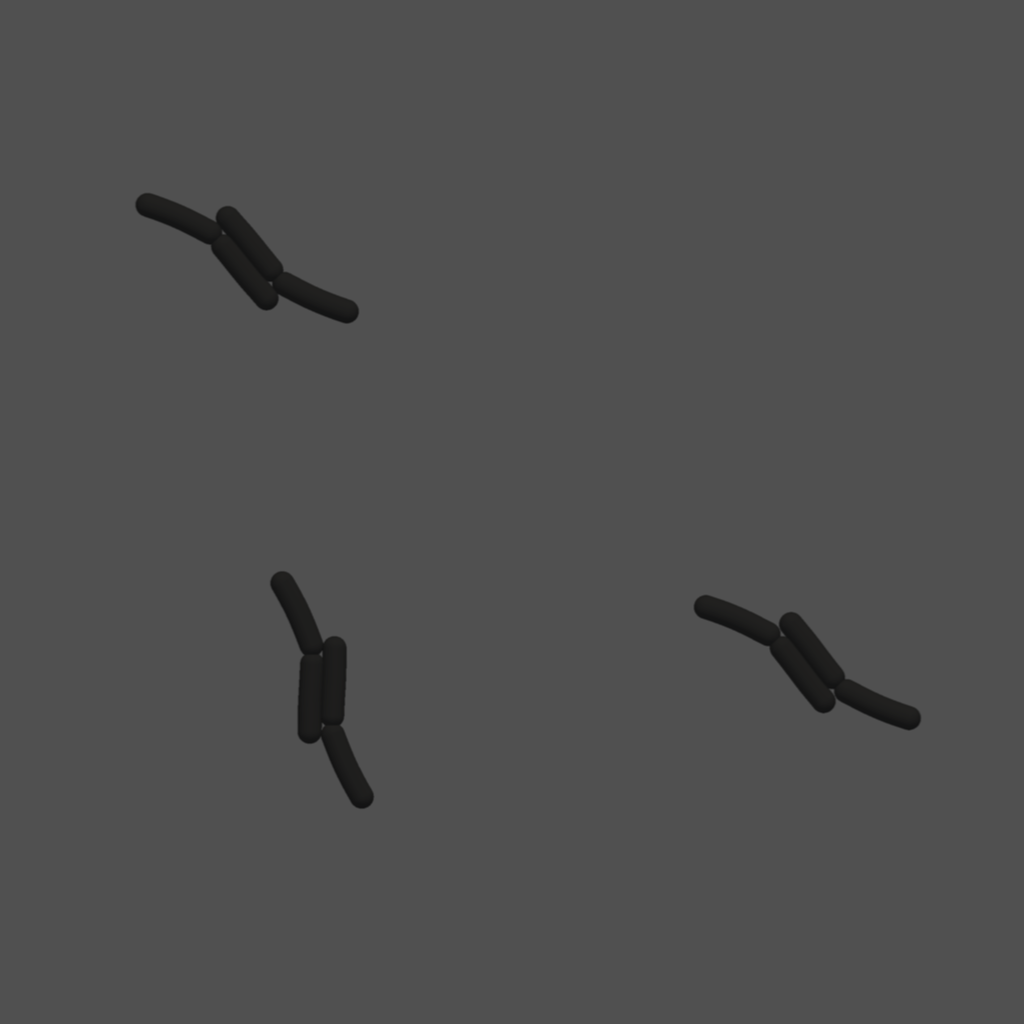
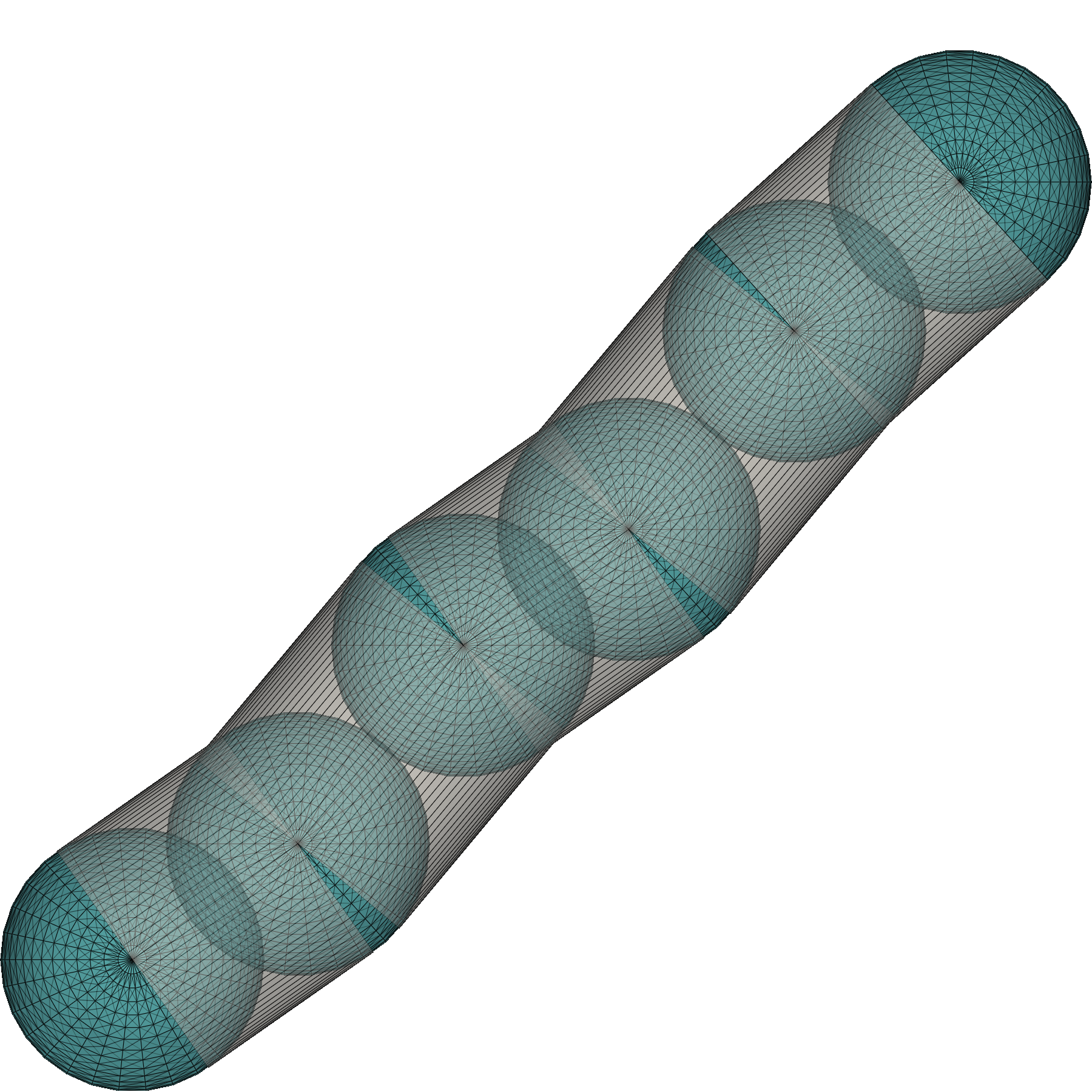


(A) Cells are rendered by combining spheres and cylinders into a single mesh. Spheres are highlighted in blue and cylinders in gray. (B) A result from combining sphere and cylinder meshes to obtain the shape of a bacterium. (C) Rendered image after applying photo-realistic filters. (D) Instance-level cell masks with unique color assignment to each agent.
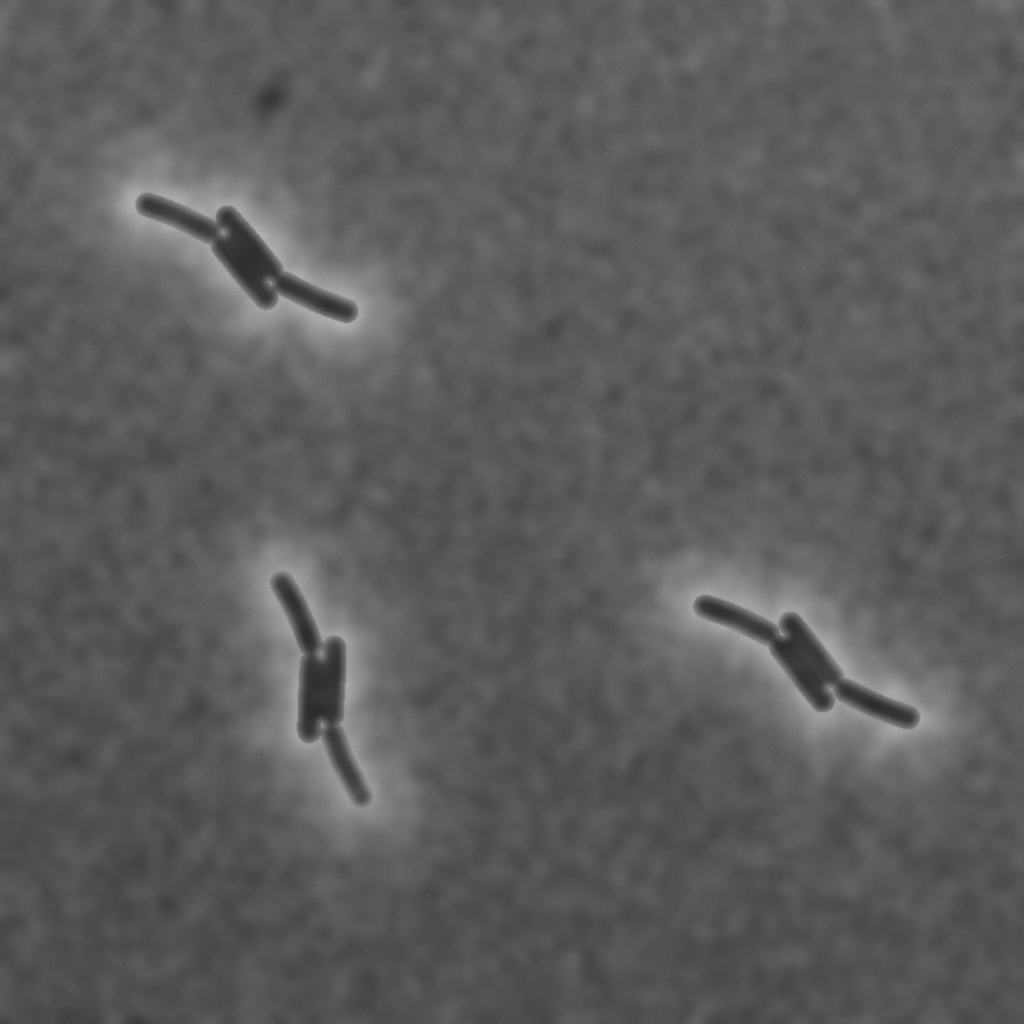
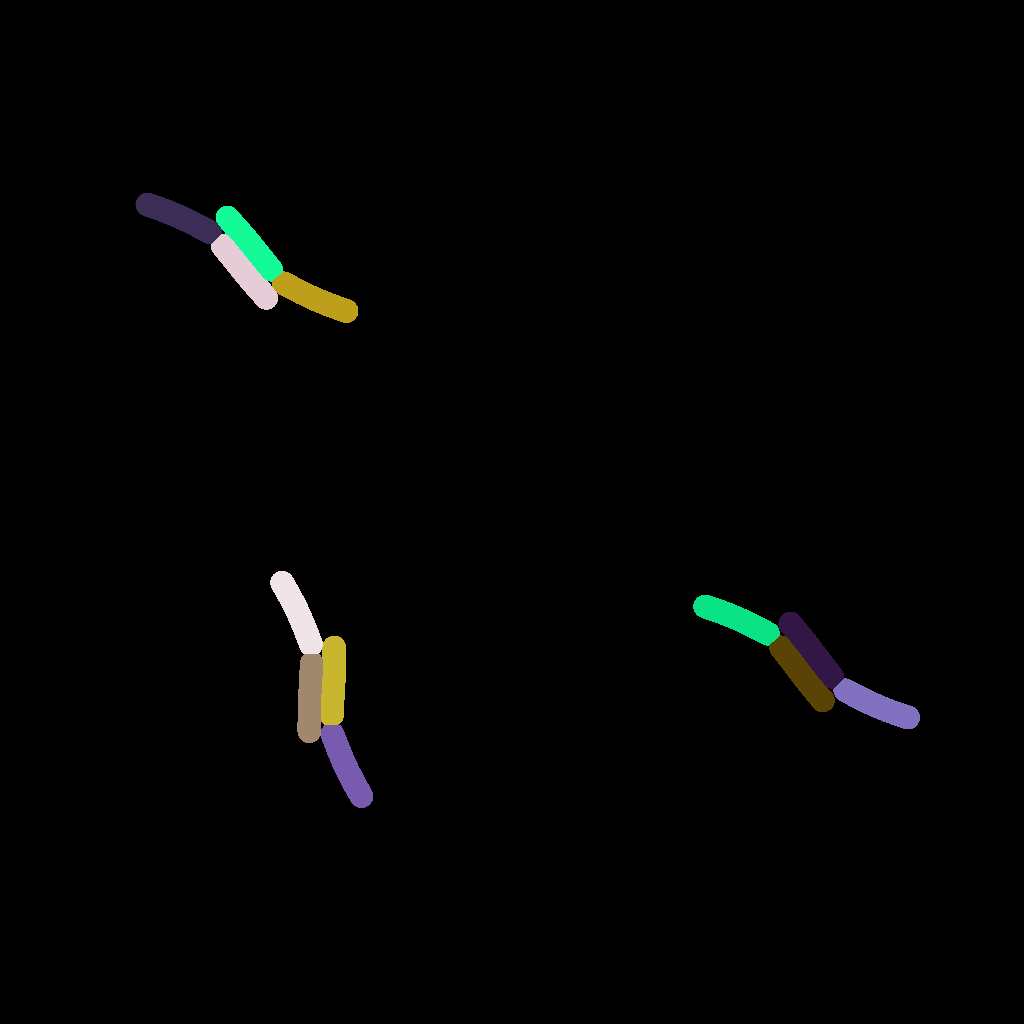
The follwing figure (A) was taken from the omnipose dataset [10] which contains a wide variety of cell types and was used to train segmentation tools.
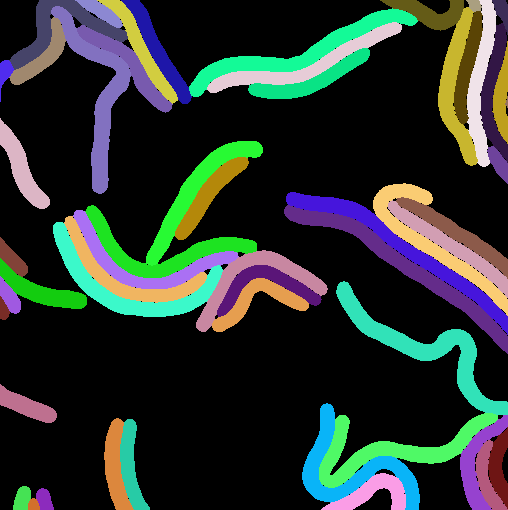
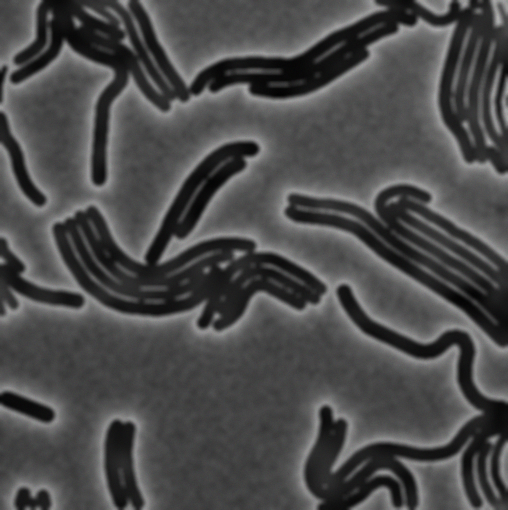



(A) Real microscopic image of the M90T strain of Shigella flexneri from the omnipose data set. (B) Synthetically generated microscopic image with optimized parameters. (C) Instance-level cell masks corresponding to (A).